from China 
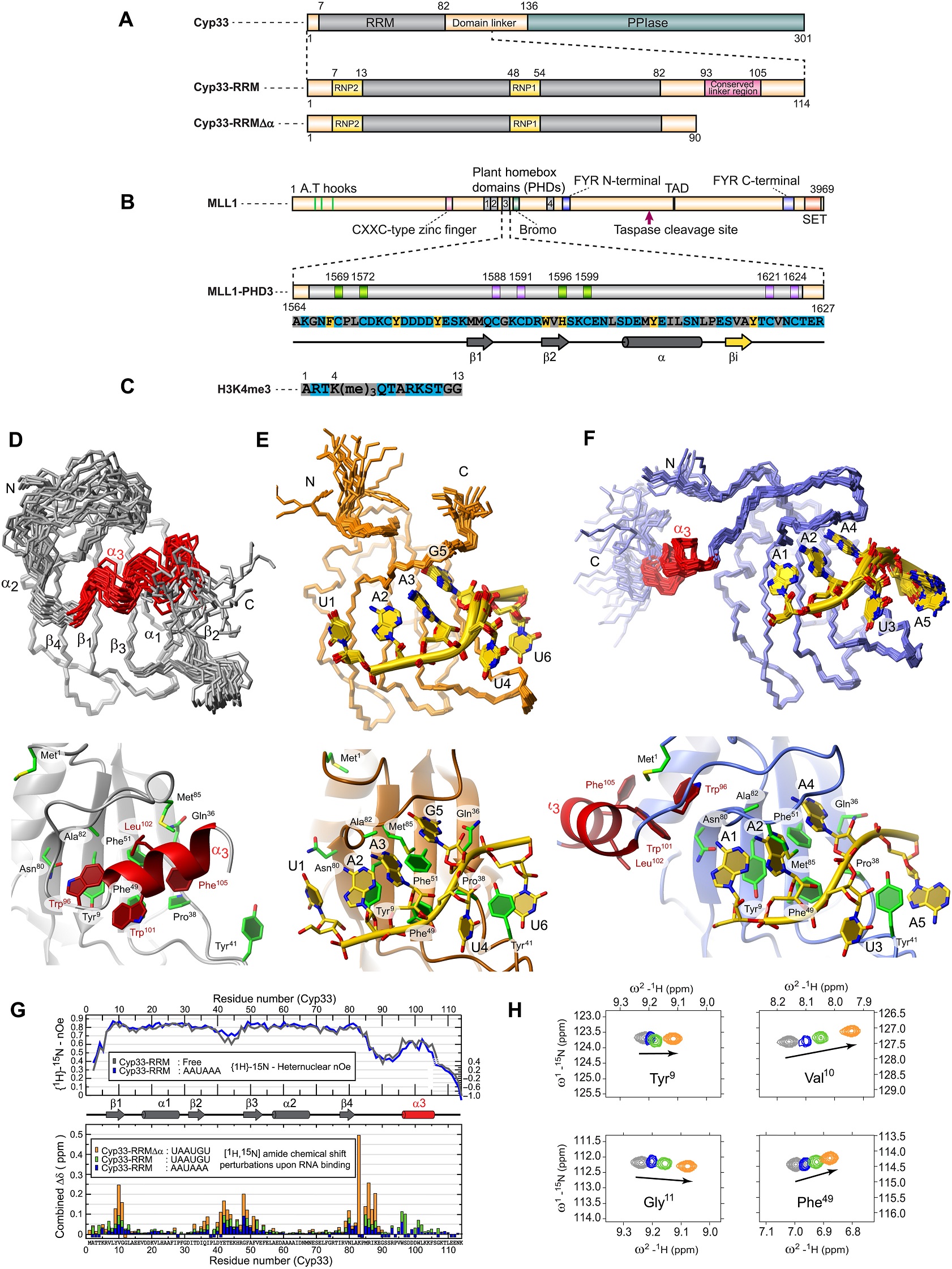
| Cyp33-RRM* | Cyp33-RRM: AAUAAA† | Cyp33-RRMΔα: UAAUGUCG‡ | Cyp33-RRM: MLL1-PHD3§ | Cyp33-RRMΔα: MLL1-PHD3:H3K4me3|| | |
|---|---|---|---|---|---|
| Completeness of 1H chem. shift assignm. (%)¶ | 96.1 | 97.2 | 99.5 | 92.4 | 89.9 |
| Cyp33 (%) | 96.1 | 96.9 | 99.4 | 92.8 | 94.9 |
| RNA or MLL1 (%) | 100 | 100 | 91.6 | 86.9 | |
| H3K4me3 (%) | 65.6 | ||||
| NMR restraints | |||||
| Distance restraints | 3112 | 3780 | 3936 | 4548 | 3262 |
| Cyp33 intramolecular | 3112 | 3627 | 3668 | 3193 | 2362 |
| Intraresidual | 562 | 643 | 576 | 552 | 454 |
| Sequential (|i − j| = 1) | 860 | 942 | 896 | 807 | 630 |
| Medium range (1 < |i − j| < 5) | 654 | 831 | 779 | 733 | 497 |
| Long range (|i − j| ≥ 5) | 1036 | 1211 | 1417 | 1101 | 781 |
| RNA or MLL1 intramolecular | 42 | 132 | 1129 | 624 | |
| Intraresidual | 36 | 98 | 246 | 175 | |
| Sequential (|i − j| = 1) | 6 | 29 | 345 | 213 | |
| Medium range (1 < |i − j| < 5) | 0 | 5 | 284 | 135 | |
| Long range (|i − j| ≥ 5) | 0 | 0 | 254 | 101 | |
| H3K4me3 intramolecular | 40 | ||||
| Intraresidual | 31 | ||||
| Sequential (|i − j| = 1) | 4 | ||||
| Medium range (1 < |i − j| < 5) | 5 | ||||
| Cyp33:RNA or Cyp33:MLL1 intermolecular | 111 | 136 | 226 | 162 | |
| MLL1:H3K4me3 intermolecular | 74 | ||||
| Torsion angles# | 0 | 60 | 8 | 176 | 0 |
| Cyp33 backbone | 0 | 54 | 0 | 122 | 0 |
| MLL1 backbone | 54 | 0 | |||
| H3K4me3 backbone | 0 | ||||
| RNA sugar pucker | 6 | 8 | |||
| Energy statistics** | |||||
| Average distance constraint violations | |||||
| 0.1–0.2 Å | 15.0 ± 2.7 | 15.0 ± 2.7 | 15.0 ± 2.7 | 9.6 ± 2.0 | 24.4 ± 4.3 |
| 0.2–0.3 Å | 0.8 ± 1.0 | 0.9 ± 1.0 | 0.1 ± 0.2 | 0.5 ± 0.5 | 4.0 ± 1.2 |
| >0.3 A | 0.6 ± 0.7 | 0.1 ± 0.2 | 0.0 ± 0.0 | 0.1 ± 0.4 | 0.2 ± 0.4 |
| Maximal (Å) | 0.27 ± 0.1 | 0.21 ± 0.03 | 0.16 ± 0.02 | 0.21 ± 0.06 | 0.28 ± 0.03 |
| Average angle constraint violations | |||||
| <5° | 22.6 ± 2.2 | 3.0 ± 0.0 | 19.6 ± 2.0 | 0.0 ± 0.0 | |
| >5° | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | |
| Maximal (°) | 1.44 ± 1.18 | 0.32 ± 0.03 | 0.64 ± 0.21 | 0.0 ± 0.0 | |
| Mean AMBER constr. viol. energy | 21.3 ± 3.0 | 24.0 ± 2.6 | 11.1 ± 0.9 | 15.4 ± 1.9 | 23.1 ± 1.7 |
| Distance | 21.3 ± 3.0 | 21.3 ± 1.6 | 10.7 ± 0.9 | 15.3 ± 1.9 | 22.9 ± 1.7 |
| Torsion | 2.8 ± 2.2 | 0.4 ± 0.1 | 0.1 ± 0.1 | 0.2 ± 0.1 | |
| Mean AMBER energy | −3174 ± 12 | −4229 ± 9 | −4099 ± 10 | −5141 ± 8 | −5261 ± 19 |
| Mean deviation from ideal covalent geometry | |||||
| Bond length (Å) | 0.004 ± 0.000 | 0.004 ± 0.000 | 0.004 ± 0.000 | 0.004 ± 0.000 | 0.004 ± 0.000 |
| Bond angle (°) | 1.651 ± 0.018 | 1.758 ± 0.012 | 1.768 ± 0.014 | 1.624 ± 0.014 | 1.641 ± 0.018 |
| Ramachandran plot statistics**,†† | |||||
| Residues in most favored regions (%) | 90.0 ± 2.1 | 82.5 ± 2.1 | 84.6 ± 1.6 | 88.2 ± 1.4 | 86.5 ± 1.9 |
| Residues in additionally allowed regions (%) | 10.0 ± 2.1 | 17.5 ± 2.1 | 15.4 ± 1.6 | 11.6 ± 1.4 | 13.0 ± 1.9 |
| Residues in generously allowed regions (%) | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.2 ± 0.4 | 0.4 ± 0.6 |
| Residues in disallowed regions (%) | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.2 | 0.2 ± 0.4 |
| RMSD to mean structure statistics** | |||||
| Cyp33 | |||||
| Backbone atoms | 0.18 ± 0.03 | 0.26 ± 0.05 | 0.12 ± 0.03 | 0.28 ± 0.04 | 0.20 ± 0.05 |
| Heavy atoms | 0.53 ± 0.08 | 0.52 ± 0.08 | 0.42 ± 0.08 | 0.59 ± 0.06 | 0.53 ± 0.04 |
| RNA or MLL1 | |||||
| Backbone atoms | 0.34 ± 0.11 | 0.19 ± 0.05 | 0.50 ± 0.11 | 0.77 ± 0.17 | |
| Heavy atoms | 0.47 ± 0.18 | 0.29 ± 0.05 | 0.84 ± 0.11 | 1.25 ± 0.20 | |
| H3K4me3 | |||||
| Backbone atoms | 0.38 ± 0.20 | ||||
| Heavy atoms | 1.03 ± 0.22 | ||||
| All molecules | |||||
| Backbone atoms | 0.18 ± 0.03 | 0.29 ± 0.05 | 0.15 ± 0.03 | 0.46 ± 0.06 | 0.63 ± 0.15 |
| Heavy atoms | 0.53 ± 0.08 | 0.54 ± 0.07 | 0.41 ± 0.07 | 0.76 ± 0.06 | 0.99 ± 0.15 |
| PDB code | 7ZEV | 7ZEW | 7ZEX | 7ZEY | 7ZEX |
| BMRB code | 34724 | 34725 | 34726 | 34727 | 34728 |